Usage
Workflow
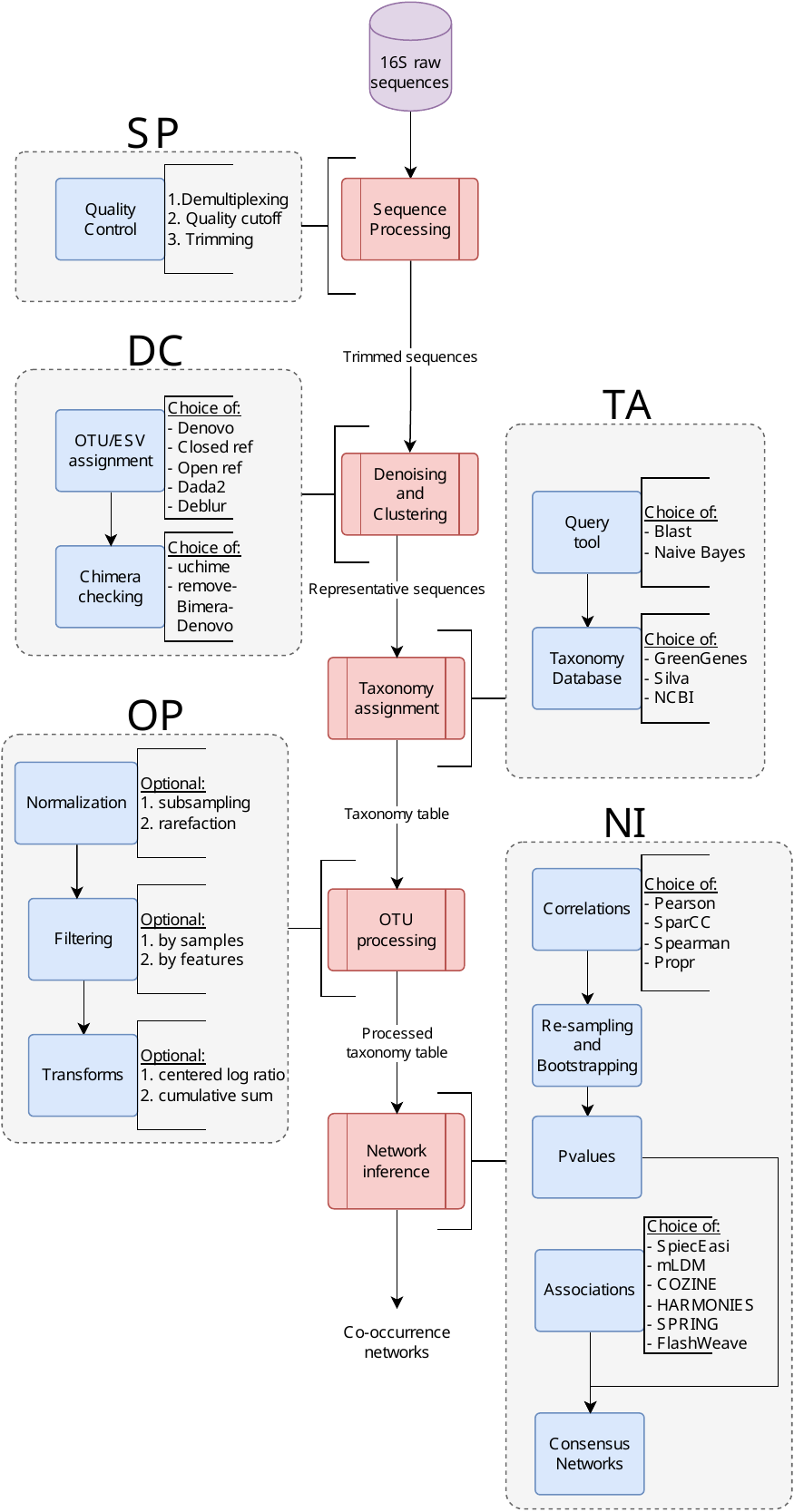
pipeline
It supports the conversion of raw 16S sequence data into co-occurrence networks. Each process in the pipeline supports alternate tools for performing the same task, users can use the configuration file to change these values.
Usage
The MiCoNE pipelines comes with an easy-to-use CLI. To get a list of
subcommands you can type:
micone --help
Supported subcommands:
install- Initializes the package and environments (createscondaenvironments for various pipeline processes)init- Initialize the nextflow templates for the micone workflowclean- Cleans files from a pipeline run (cleans temporary data, log files and other extraneous files)validate-results- Check the results of the pipeline execution
Installing the environments
In order to run the pipeline various conda environments must first
be installed on the system. Use the following comand to initialize all
the environments:
micone install
Or to initialize a particular environment use:
micone install -e "micone-qiime2"
The list of supported environments:
micone-cozine
micone-dada2
micone-flashweave
micone-harmonies
micone-mldm
micone-propr
micone-qiime2
micone-sparcc
micone-spieceasi
micone-spring
Initializing the pipeline template
To initialize the full pipeline (from raw 16S sequencing reads to co-occurrence networks):
micone init -w <workflow> -o <path/to/folder>
Other supported pipeline templates are (work in progress):
full
ni
op_ni
ta_op_ni
To run the pipeline, update the relevant config files (see next
section), activate the micone environment and run the run.sh
script that was copied to the directory:
bash run.sh
This runs the pipeline locally using the config options specified.
To run the pipeline on an SGE enabled cluster, add the relevant
project/resource allocation flags to the run.sh script and run as:
qsub run.sh
Configuration and the pipeline template
The pipeline template for the micone “workflow” (see previous section
for list of supported options) is copied to the desired folder after
running micone init -w <workflow>. The template folder contains the
following folders and files:
nf_micone: Folder contatining the
miconedefault configs, data, functions, and modulestemplates: Folder containing the templates (scripts) that are executed during the pipeline run
main.nf: The pipeline “workflow” defined in the
nextflowDSL 2 specificationnextflow.config: The configuration for the pipeline. This file needs to be modified in order to change any configuration options for the pipeline run
metadata.json: Contains the basic metadata that describes the dataset that is to be processed. Should be updated accordingly before pipeline execution
samplesheet.csv: The file that contains the locations of the input data necessary for the pipeline run. Should be updated accordingly before pipeline execution
run.sh: The
bashscript that contains commands used to execute thenextflowpipeline
The folder nf_micone/configs contains the default configs for all
the micone pipeline workflows. These options can also be viewed in
tabular format in the
documentation.
For example, to change the tool used for OTU assignment to dada2 and
deblur, you can add the following to nextflow.config:
// ... config initialization
params {
// ... other config options
denoise_cluster {
otu_assignment {
selection = ['dada2', 'deblur']
}
}
}
Example configuration files used for the analyses in the manuscript can be found here.
Visualization of results (coming soon)
The results of the pipeline execution can be visualized using the scripts in the manuscript repo
Configuring the pipeline
The parameters for the pipeline execution are in the micone/pipelines/configs/*.config in the MiCoNE GitHub repository. These can be configured using the nextflow.config file.
The following tables contain the list of default parameters for each step of the pipeline:
Sequence Processing (SP)
Step |
Task |
Tool |
Parameter |
Value |
|---|---|---|---|---|
Sequence Processing |
Demultiplexing |
demultiplexing_illumina_single |
barcode_column |
barcode-sequence |
Sequence Processing |
Demultiplexing |
demultiplexing_illumina_single |
rev_comp_barcodes |
false |
Sequence Processing |
Demultiplexing |
demultiplexing_illumina_single |
rev_comp_mapping_barcodes |
false |
Sequence Processing |
Demultiplexing |
demultiplexing_illumina_paired |
barcode_column |
barcode-sequence |
Sequence Processing |
Demultiplexing |
demultiplexing_illumina_paired |
rev_comp_barcodes |
false |
Sequence Processing |
Demultiplexing |
demultiplexing_illumina_paired |
rev_comp_mapping_barcodes |
false |
Sequence Processing |
Trimming |
export_visualization_single |
seq_samplesize |
10000 |
Sequence Processing |
Trimming |
export_visualization_paired |
seq_samplesize |
10000 |
Sequence Processing |
Trimming |
trimming_single |
ncpus |
1 |
Sequence Processing |
Trimming |
trimming_single |
max_ee |
2 |
Sequence Processing |
Trimming |
trimming_single |
trunc_q |
2 |
Sequence Processing |
Trimming |
trimming_paired |
ncpus |
1 |
Sequence Processing |
Trimming |
trimming_paired |
max_ee |
2 |
Sequence Processing |
Trimming |
trimming_paired |
trunc_q |
2 |
Denoising and Clustering (DC)
Task |
Tool |
Parameter |
Value |
|---|---|---|---|
OTU assignment |
Closed reference |
ncpus |
1 |
OTU assignment |
Closed reference |
percent_identity |
0.97 |
OTU assignment |
Closed reference |
strand |
“plus” |
OTU assignment |
Closed reference |
reference_sequences |
“gg_97” |
OTU assignment |
Open reference |
ncpus |
1 |
OTU assignment |
Open reference |
percent_identity |
0.97 |
OTU assignment |
Open reference |
strand |
“plus” |
OTU assignment |
Open reference |
reference_sequences |
“gg_97” |
OTU assignment |
De novo |
ncpus |
1 |
OTU assignment |
De novo |
percent_identity |
0.97 |
OTU assignment |
Dada2 |
ncpus |
1 |
OTU assignment |
Dada2 |
big_data |
“FALSE” |
OTU assignment |
Deblur |
ncpus |
1 |
OTU assignment |
Deblur |
min_reads |
2 |
OTU assignment |
Deblur |
min_size |
2 |
Chimera checking |
Remove bimera |
ncpus |
1 |
Chimera checking |
Remove bimera |
chimera_method |
“consensus” |
Taxonomy Assignment (TA)
Task |
Tool |
Parameter |
Value |
|---|---|---|---|
Assign |
Naive Bayes |
classifer |
“gg_13_8_99_515_806” |
Assign |
Naive Bayes |
confidence |
0.7 |
Assign |
Naive Bayes |
ncpus |
1 |
Assign |
BLAST |
references |
“ncbi_refseq” |
Assign |
BLAST |
max_accepts |
10 |
Assign |
BLAST |
perc_identity |
0.8 |
Assign |
BLAST |
evalue |
0.001 |
Assign |
BLAST |
min_consensus |
0.51 |
OTU Processing (OP)
Task |
Tool |
Parameter |
Value |
|---|---|---|---|
Transform |
Fork |
axis |
“sample” |
Transform |
Fork |
column |
“” |
Transform |
Normalize & Filter |
axis |
“None” |
Transform |
Normalize & Filter |
count_thres |
500 |
Transform |
Normalize & Filter |
prevalence_thres |
0.05 |
Transform |
Normalize & Filter |
obssum_thres |
100 |
Transform |
Normalize & Filter |
rm_sparse_obs |
“[true, false]” |
Transform |
Normalize & Filter |
rm_sparse_samples |
true |
Transform |
Normalize & Filter |
abundance_thres |
0.01 |
Transform |
Group |
tax_levels |
“[‘Phylum’, ‘Class’, ‘Order’, ‘Family’, ‘Genus’, ‘Species’]” |
Network Inference (NI)
Task |
Tool |
Parameter |
Value |
|---|---|---|---|
Bootstrap |
Resample |
bootstraps |
1000 |
Bootstrap |
Resample |
ncpus |
1 |
Bootstrap |
Pvalue |
slim |
false |
Bootstrap |
Pvalue |
ncpus |
1 |
Correlation |
sparcc |
ncpus |
1 |
Correlation |
sparcc |
iterations |
50 |
Correlation |
pearson |
ncpus |
1 |
Correlation |
spearman |
ncpus |
1 |
Correlation |
propr |
ncpus |
1 |
Direct |
spieceasi |
method |
“mb” |
Direct |
spieceasi |
ncpus |
1 |
Direct |
spieceasi |
nreps |
50 |
Direct |
spieceasi |
nlambda |
20 |
Direct |
spieceasi |
lambda_min_ratio |
1e-2 |
Direct |
flashweave |
ncpus |
1 |
Direct |
flashweave |
sensitive |
“true” |
Direct |
flashweave |
heterogeneous |
“false” |
Direct |
flashweave |
fdr_correction |
“true” |
Direct |
mldm |
Z_mean |
1 |
Direct |
mldm |
max_iteration |
1500 |
Direct |
cozine |
lambda_min_ratio |
0.1 |
Direct |
harmonies |
iterations |
10000 |
Direct |
harmonies |
sparsity_cutoff |
0.5 |
Direct |
spring |
ncpus |
1 |
Direct |
spring |
nlambda |
20 |
Direct |
spring |
lambda_min_ratio |
0.01 |
Network |
Make network with pvalues |
interaction_threshold |
0.3 |
Network |
Make network with pvalues |
pvalue_threshold |
0.05 |
Network |
Make network with pvalues |
metadata_file |
“” |
Network |
Make network without pvalues |
interaction_threshold |
0.3 |
Network |
Make network without pvalues |
metadata_file |
“” |
Network |
Merge pvalues |
id_field |
“taxid” |
Network |
Create consensus |
method |
“scaled_sum” |
Network |
Create consensus |
parameter |
0.333 |
Network |
Create consensus |
pvalue_filter |
“true” |
Network |
Create consensus |
interaction_filter |
“true” |
Network |
Create consensus |
id_field |
“taxid” |